COMPOUND PROPERTY CALCULATION & REMOVAL OF UNDESIRABLES
FILTER
FILTER is a very fast molecular filtering and selection application. It uses a combination of physical property calculations and functional group knowledge to remove undesirable compounds before they enter experimental or virtual screening.
Undesirable properties may include: toxic functionalities, a high likelihood of binding covalently with the target protein, interfering with the experimental assay, and/or a low probability of oral bioavailability.
Eliminating unwanted molecules before the use of modeling applications will, in turn, substantially increase the positive predictive value of these tools, and significantly reduce their processing time.
For more detailed information on FILTER, please click below:
Documentation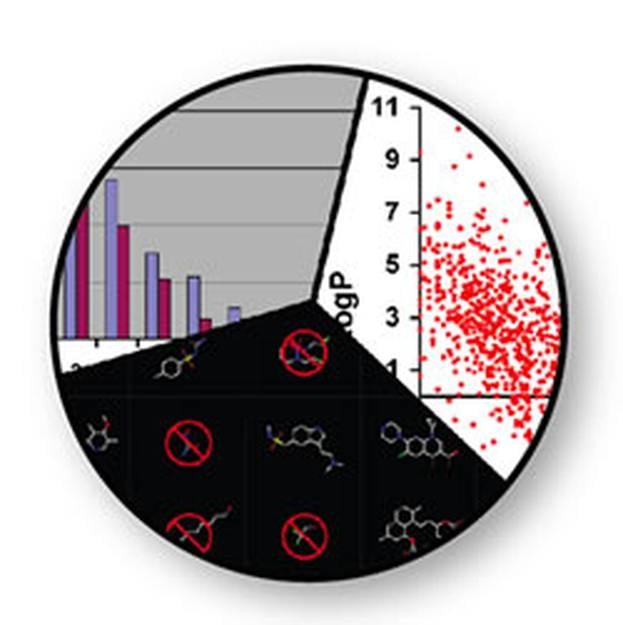
Removing undesirable compounds early makes the downstream processes significantly more efficient; a simple concept that is too often ignored.
Features
- Uses predefined filter files - text files that can be easily customized
- Filters based upon calculated properties such as: MW, XlogP [1], XlogS, PSA [2], hydrogen bond donor and acceptor count, rotatable bonds, ring size and number, etc.
- Removes or retains molecules based upon pre-defined or user-defined substructures
- Assigns graph-based protonation state for consistency and speed
- Offers ADME filters such as Lipinski [3], Egan [4], Veber [5] and Martin [6]
- Removes compounds with impossible bonding and inappropriate elements
- Generates tab-separated files suitable for import into spreadsheets
- Processes 400 mol/sec
References
- A New Atom-Additive Method for Calculating Partition Coefficients Wang, R., Ying, F., and Lai, L., J. Chem. Inf. Comput. Sci., 1997, 37, 615.
- Fast calculation of molecular polar surface area as a sum of fragment-based contributions and its application to the prediction of drug transport properties Ertl, P., Rohde, B., and Selzer, P., J. Med. Chem., 2000, 43, 3714.
- Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings Lipinski, C., et al., Adv. Drug Deliv. Rev., 1997, 23, 3.
- Prediction of drug absorption using multivariate statistics Egan, W.J., Merz, K.M., Baldwin, J.J., J. Med. Chem., 2000, 43, 3867.
- Molecular Properties That Influence the Oral Bioavailability of Drug Candidates Veber, D.F., Johnson, S.R., Cheng, H.Y., Smith, B.R., Ward, K.W., Kipple, K.D., J. Med. Chem., 2002, 45, 2615.
- A bioavailability score Martin, Y.C., J. Med. Chem., 2005, 48, 3164.
Webinar: Target X: An Unobstructed View of Pockets
Webinar: Own Your Own Target with Target X
Webinar: Modular Molecular Modeling
Webinar: ML-Enabled integration of affinity prediction and lead discovery: 3D-QSAR
Webinar: Novel Hits from Beyond the Known: ROCS X
Webinar: Exploring the Uncharted: Discovery at Trillion-Scale with ROCS X
Webinar: Own Your Own Target with Target X
Webinar: Modular Molecular Modeling
Webinar: ML-Enabled integration of affinity prediction and lead discovery: 3D-QSAR
Webinar: Novel Hits from Beyond the Known: ROCS X
Webinar: Exploring the Uncharted: Discovery at Trillion-Scale with ROCS X
Resources
View Our Recent Webinars
Upcoming Webinar
Webinar: Target X: An Unobstructed View of Pockets
Webinar: Target X: An Unobstructed View of Pockets About this session
Register now
Blog Post
Webinar: Own Your Own Target with Target X
Webinar: Own Your Own Target with Target X Join us on Thursday, March 19th to hear fromDavid Lebard, Head of Target Exploration, at OpenEye, Cadence Molecular Sciences, about OpenEye's revolutionary Pocket Detection Method.
Read now


