Target X: Pocket Detection and Ligandability Assessment Model
The identification and exploitation of binding sites in protein structures often represents a pivotal step in designing effective therapeutics. Sites that are rarely or never observed in experimental structures, yet remain druggable, are referred to as cryptic pockets. These pockets offer novel opportunities for modulating the activity of a target protein and are particularly valuable in crafting isoform-selective ligands when the structure of the substrate-binding site remains consistent across different variants of the target protein.
Through the use of state-of-the-art enhanced sampling molecular dynamics simulations, OpenEye's Target X pocket detection tools and ligandability assessment model empower users to thoroughly explore a protein's conformational space, potentially revealing one or more cryptic pockets. Customized visual and quantitative analysis of the results can yield valuable insights into the presence and potential druggability of these pockets.
Contact us to learn how you can gain new therapeutic opportunities using OpenEye's Target X.

Features
-
Performance. Fast and efficient computation using Weighted Ensemble MD for efficient exploration of potential binding sites.
-
Flexible. Handles single and mixed solvent simulations.
- Deeper Insight. Start your drug discovery campaign with deeper structural insight and reduce reliance on a single binding-site strategy.
- Detect Cryptic Pockets. Multiple pocket detection method options provide versatility for researchers.
- Rank Pocket Ligandability. A reliable ligandability prediction model helps prioritize protein pockets and streamline their use for scientists.
-
Turn-key. Automated single step end-to-end workflows.
Uncover hidden features in protein structures
Transitioning from static structures to dynamic insights is a critical aspect of computational chemistry.
OpenEye provides users with automated workflows to explore hidden putative ligand binding sites for challenging protein targets.
With Target X, users can:
- Prepare and execute Weighted Ensemble Molecular Dynamics (WE-MD) simulations in a single or mixed solvent.
- Conduct pocket detection analysis to identify potential cryptic pocket sites.
- Rank pockets identified from molecular dynamics simulations, or directly on static protein structures using the ligandability model.
With OpenEye's Target X on the Orion® Molecular Design Platform, scientists can run calculations across hundreds or even thousands of GPUs in the cloud, saving valuable discovery time.


Empowering Structural Biology
How do dynamics impact drug efficacy and resistance? Target X helps structural biologists look beyond static protein structures by revealing transient or hidden binding pockets that may not appear in apo or holo crystal structures. By simulating natural protein motions and ranking newly found sites based on predicted ligandability, Target X provides a systematic way to evaluate potential allosteric sites, design protein variants to stabilize relevant conformations, and plan follow-up experiments such as mutagenesis or fragment screening. With Target X, you can effectively translate your structural findings into therapeutic strategies.
Start Your Drug Discovery Campaign with an Advantage
Why run Target X on all your drug discovery targets? Target X unlocks the full conformational landscape of your protein, allowing you to start your drug discovery campaign with deeper structural insight. This can open parallel discovery paths and reduce reliance on a single binding-site strategy.
Target X provides an early, molecular-level view of how your protein truly behaves using a rigorous, physics-based approach. By combining enhanced conformational sampling with a reliable ligandability prediction model, it reveals transient and cryptic pockets that static structures often miss. Target X enables structural biology and chemistry teams to prioritize strategies with deeper insight before committing experimental resources.

Discover new therapeutic opportunities
OpenEye’s Target X enables scientists to detect cryptic pockets, uncover alternative binding sites not apparent from a protein’s primary function or known active sites, and reliably rank pockets using ligandability model. Identifying such hidden pockets can open new therapeutic opportunities and support the design of novel drugs.
-
OpenEye’s Target X comes with fully automated workflows to simplify your calculations. Users can access an end-to-end workflow that automated these steps:
-
Solvate and equilibrate target proteins (in single or mixed solvent).
-
Calculate normal modes.
-
Perform a Weighted Ensemble MD simulation.
-
Run cryptic pocket detection search.
- Assess and rank pocket ligandabilty.
-
-
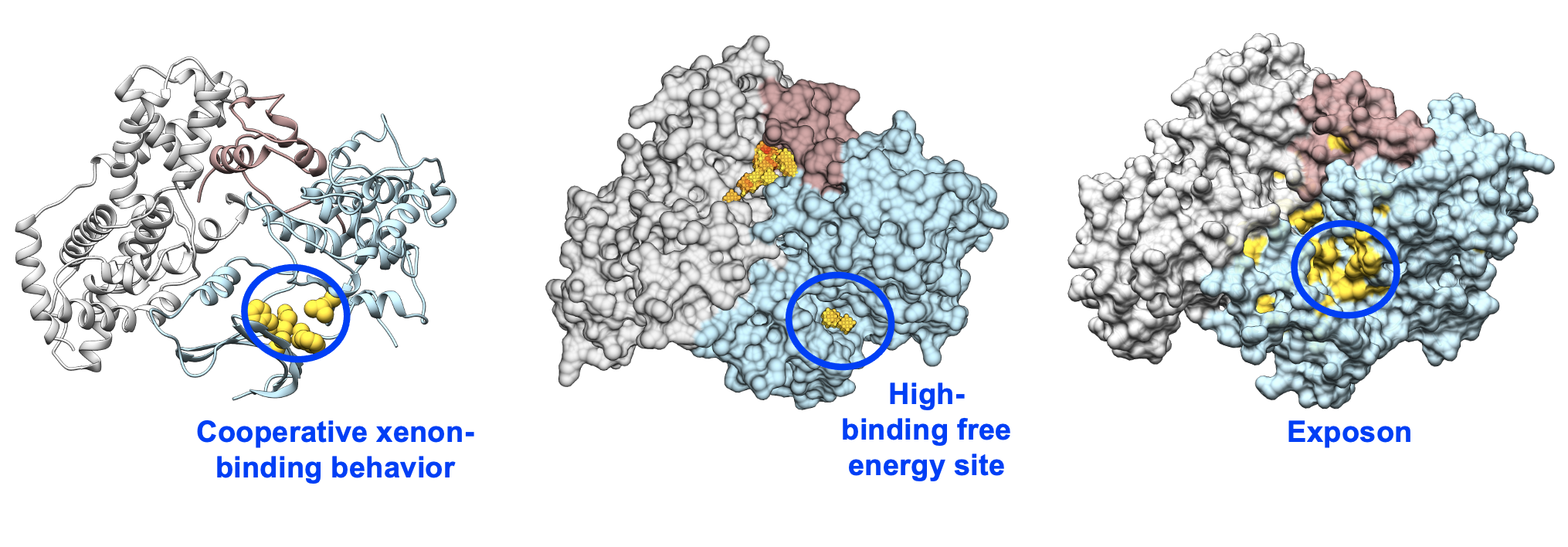
It's important to note that no single pocket detection method is foolproof, and a combination of different approaches may be used to increase the accuracy and reliability of detecting cryptic pockets in proteins. Hence, OpenEye provides three independent methods for your cryptic pocket detection:
- Exposon Analysis (Solvent accessible surface area change, single solvent only)
- CoSolvent Binding Free Energy Analysis (Probe-map analysis)
- Cooperative CoSolvent Binding Analysis (Dynamic probe binding analysis)
-
With the OpenEye’s Cryptic Pocket method, users can perform protein sampling simulations that are solvated in water (single solvent) and/or are solvated in water and xenon (mixed solvent).
-
OpenEye’s Target X method offers the advantage of using water and xenon (mixed solvent) in the protein sampling. Using xenon as a probe offers advantages including:
- Xenon is a non-selective binder to hydrophobic sites1
- Xenon has a fast diffusion rate2
- Xenon localization has been observed in pocket composed of hydrophobic and hydrophilic residues1
- Schiltz M et al., Structure. 1995, 3, 309
- Zhao Z et al., Biophys J. 2022 Dec 6;121(23):4635
-
OpenEye’s Target X offers a cost-effective approach to uncover binding sites. Compute costs may vary depending on the protein size, but the runs typically average a few hundred dollars.
Learn More
OpenEye's Target X cryptic pocket detection tools and ligandability assessment model can help you minimize the risk of off-target activity and undesired side effects, and thus save costs by reducing failure rates.
Download OpenEye Science Brief on Target X - Unlocking Hidden Binding Sites with Advanced Protein Dynamics.
Download OpenEye Science Brief on Target X - Reliable Protein Pocket Ligandability Prediction.
Watch OpenEye's recorded webinar from March 2026 for Dr. David LeBard's presentation on Own Your Own Target with Target X: Expand pocket space, prioritize by ligandability, and de-risk drug discovery decisions.
Watch OpenEye's recorded webinar from August 2025 for Dr. David LeBard's presentation on Drugging the undruggable: A highly accurate method for detecting and ranking cryptic pocket.
References
- Target X built on Groovy: An unbiased and robust ligandability prediction model built and evaluated on non-redundant, well-curated dataset. Research Square. Preprint. Feb 12, 2026. Neha Vithani, David Wych, She Zhang, Phu Tang, Alex Demidov, A. Geoffrey Skillman, David N. LeBard. DOI: https://doi.org/10.21203/rs.3.rs-8833408/v1
- Enhanced sampling and ligandability assessment to expand the repertoire of potentially druggable cryptic pockets. Research Square. Preprint. Feb 15, 2026. Neha Vithani, She Zhang, Judith Günther, Hans Purkey, J. David Lawson, Anthony Nicholls, A. Geoffrey Skillman, and David N. LeBard. DOI: https://doi.org/10.21203/rs.3.rs-8854093/v1
- Lessons Learned from a Ligand-Unbinding Stress Test for Weighted Ensemble Simulations. Bogetti AT, Yang DT, Piston HE, LeBard DN, Chong LT. ACS Omega. 2025 Jun 16;10(25):27617-27624. doi: 10.1021/acsomega.5c03809. PMID: 40620993; PMCID: PMC12223804.
- Reducing weighted ensemble variance with optimal trajectory management. Ryu WH, Russo JD, Johnson MS, Copperman JT, Thompson JP, LeBard DN, Webber RJ, Simpson G, Aristoff D, Zuckerman DM. J Chem Phys. 2026 Mar 7;164(9):094110. doi: 10.1063/5.0311015. PMID: 41773795; PMCID: PMC12959359.
- Exploration of Cryptic Pockets Using Enhanced Sampling Along Normal Modes: A Case Study of KRAS G12D. Vithani N, Zhang S, Thompson JP, Patel LA, Demidov A, Xia J, Balaeff A, Mentes A, Arnautova YA, Kohlmann A, Lawson JD, Nicholls A, Skillman AG, LeBard DN. J Chem Inf Model. 2024 Nov 11;64(21):8258-8273. doi: 10.1021/acs.jcim.4c01435. Epub 2024 Oct 17. PMID: 39419500; PMCID: PMC11558672.
- Cooperative Changes in Solvent Exposure Identify Cryptic Pockets, Switches, and Allosteric Coupling. Porter, J. R.; Moeder K. E.; Sibbald C. A.; Zimmerman M. I.; Hart K. M.; Greenberg M. J.; Bowman G. R., Biophys J. 2019, 116(5), 818-830. doi: 10.1016/j.bpj.2018.11.3144. Epub 2019 Jan 25. PMID: 30744991; PMCID: PMC6400826.
Webinar: Own Your Own Target with Target X
Webinar: Modular Molecular Modeling
Webinar: ML-Enabled integration of affinity prediction and lead discovery: 3D-QSAR
Webinar: Novel Hits from Beyond the Known: ROCS X
Webinar: Exploring the Uncharted: Discovery at Trillion-Scale with ROCS X
Resources
View Our Recent Webinars
Upcoming Webinar
Webinar: Target X: An Unobstructed View of Pockets
On Demand Webinars
Webinar: Own Your Own Target with Target X


