Trusted Science. Delivered the Way You Need.
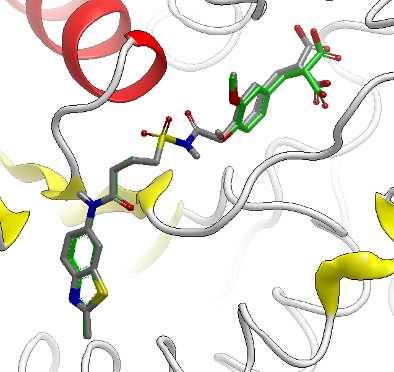
Structure-Based Design
Take your structure-centric discovery efforts to the next level with solutions that address the unique challenges of studying protein-ligand interactions. Whether you’re looking to dock and score candidates, compare protein binding sites, or estimate free-energy binding affinities, you can trust that OpenEye methods are validated, robust, and scalable.
Trusted Science.
Binding Free Energy Calculation (FE-NES)
Ultra Large-Scale Virtual Screening (Gigadock™, Gigadock Warp)
Docking (FRED, HYBRID), Posing (POSIT) and Induced-Fit Posing
Rapid and Accurate Conformers Generation (OMEGA)
Fragment Replacement for Molecular Design (BROOD)
Ligand Optimization Based on Nearby Water Energetics (SZMAP/Gameplan)
Force Field Ligand Optimization With and Without Solvent Effects (SZYBKI/Freeform)
Delivered the Way You Need.
Webinar: Target X: An Unobstructed View of Pockets
Webinar: Own Your Own Target with Target X
Webinar: Modular Molecular Modeling
Webinar: ML-Enabled integration of affinity prediction and lead discovery: 3D-QSAR
Webinar: Novel Hits from Beyond the Known: ROCS X
Webinar: Exploring the Uncharted: Discovery at Trillion-Scale with ROCS X
Webinar: Own Your Own Target with Target X
Webinar: Modular Molecular Modeling
Webinar: ML-Enabled integration of affinity prediction and lead discovery: 3D-QSAR
Webinar: Novel Hits from Beyond the Known: ROCS X
Webinar: Exploring the Uncharted: Discovery at Trillion-Scale with ROCS X
Resources
View Our Recent Webinars
Upcoming Webinar
Webinar: Target X: An Unobstructed View of Pockets
Webinar: Target X: An Unobstructed View of Pockets About this session
Register now
Blog Post
Webinar: Own Your Own Target with Target X
Webinar: Own Your Own Target with Target X Join us on Thursday, March 19th to hear fromDavid Lebard, Head of Target Exploration, at OpenEye, Cadence Molecular Sciences, about OpenEye's revolutionary Pocket Detection Method.
Read now


