OpenEye Solutions
SCIENCE | SPEED | SCALE
Trusted Science. Delivered the Way You Need.
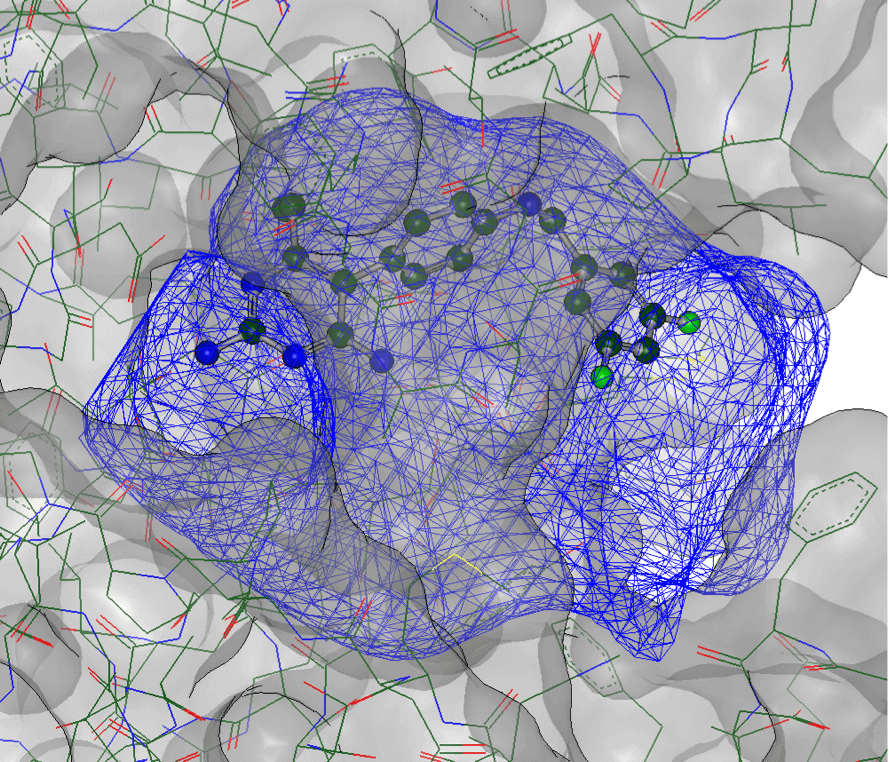
Large Scale Virtual Screening
Search billions of molecules across different chemical libraries. OpenEye delivers 2D and 3D virtual screening solutions at the scale you need, the speed you want, and the computing cost you desire.
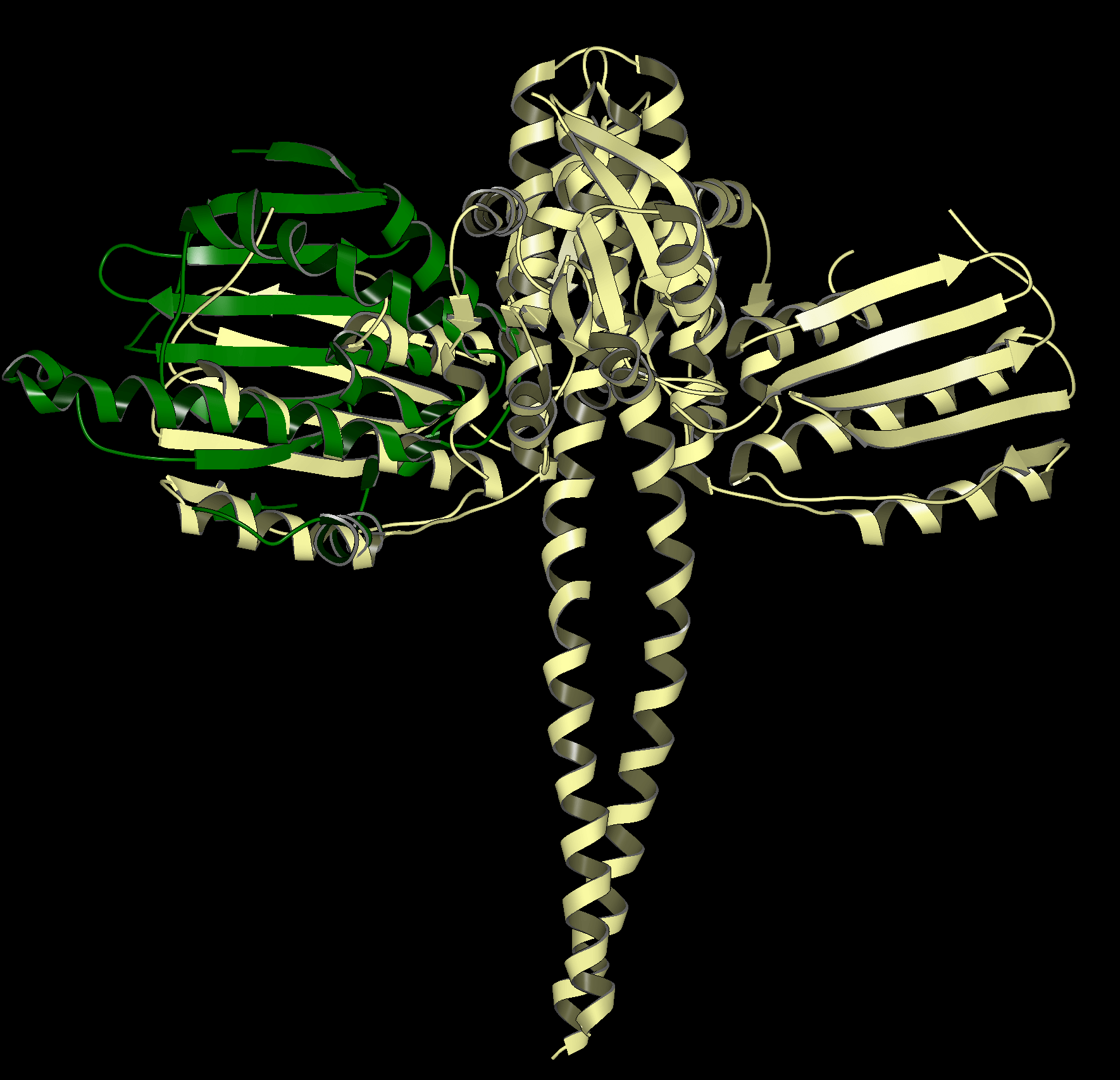
Structural Biology
Get more accurate results with careful preparation of experimental structures for downstream modeling applications. OpenEye provides automated tools that can generate high quality models, understand protein motions, uncover cryptic pockets, and streamline your protein preparation process.
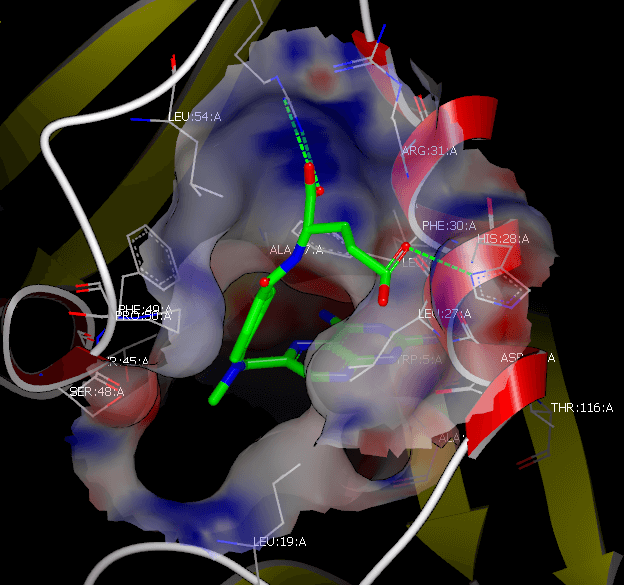
Lead Discovery
Rapidly discover new, biologically active chemical matter as potential leads in the drug discovery programs. Scientists trust OpenEye solutions for lead identification that are proven, fast and reliable.
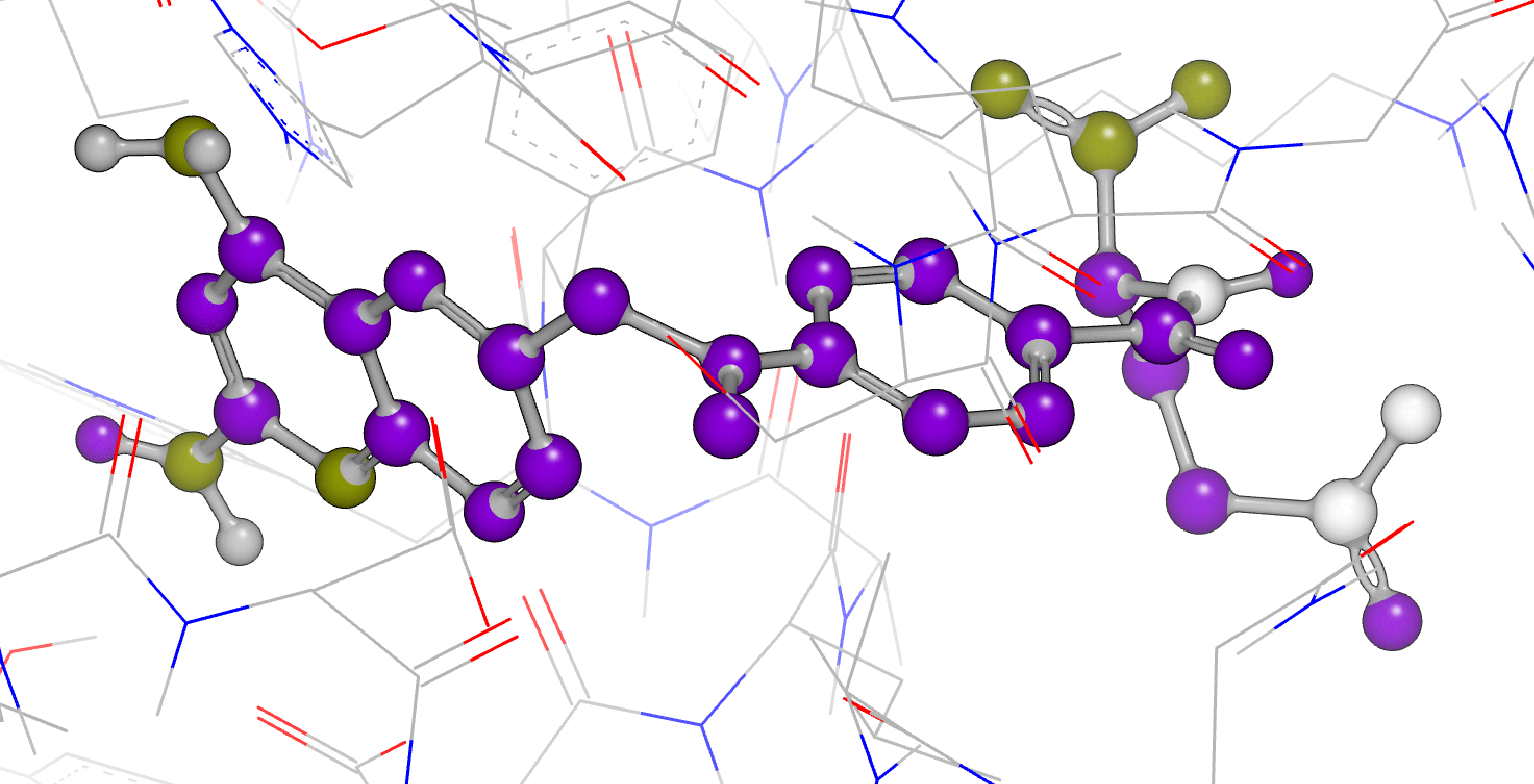
Lead Optimization
Fine-tune profiles of your preclinical lead candidates for higher likelihood of a successful progression into clinical development. OpenEye provides a wide range of cutting-edge science to optimize both the biological activity and the properties of your lead series.
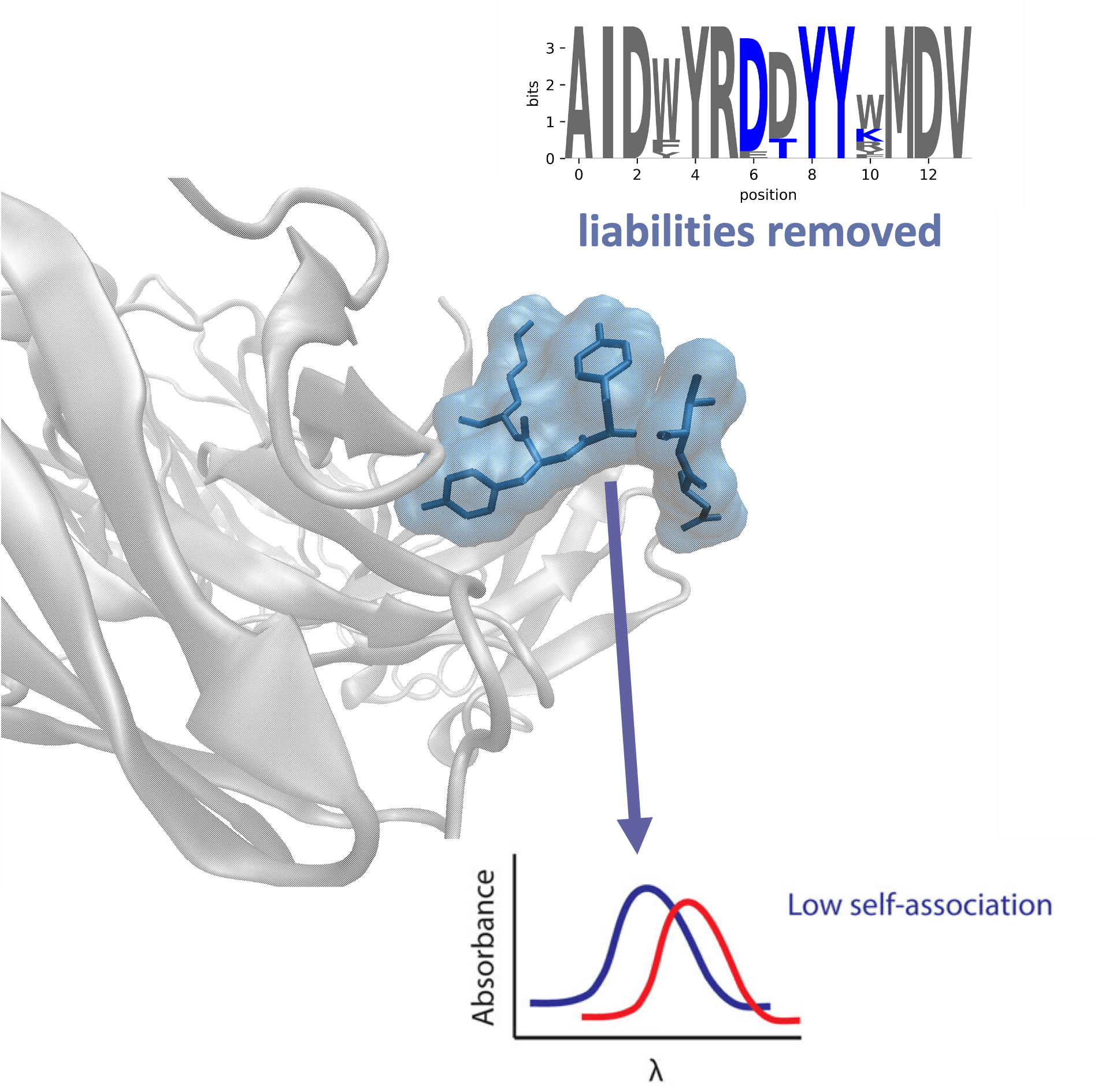
Antibody Discovery
Rapidly analyze your antibody-based sequencing dataset with cloud computing. OpenEye provides you the ability to process tens of millions of sequences from the convenience of your web browser. Increase the number of your clonotype leads 5-10x compared to random colony screening.
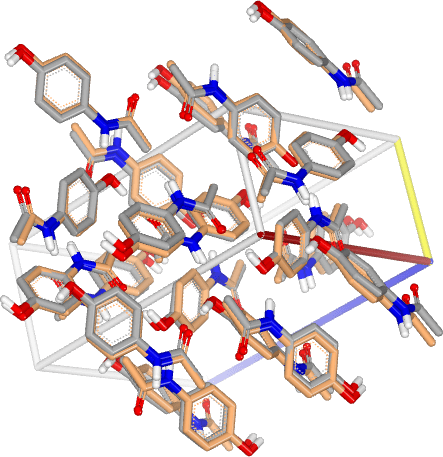
Drug Formulation
Leverage computational advances to cost-effectively predict downstream development profile of your drug leads. From pharmaceutical crystals predictions to intrinsic properties calculations, OpenEye provides you with the necessary tools to get ahead of your competitors.
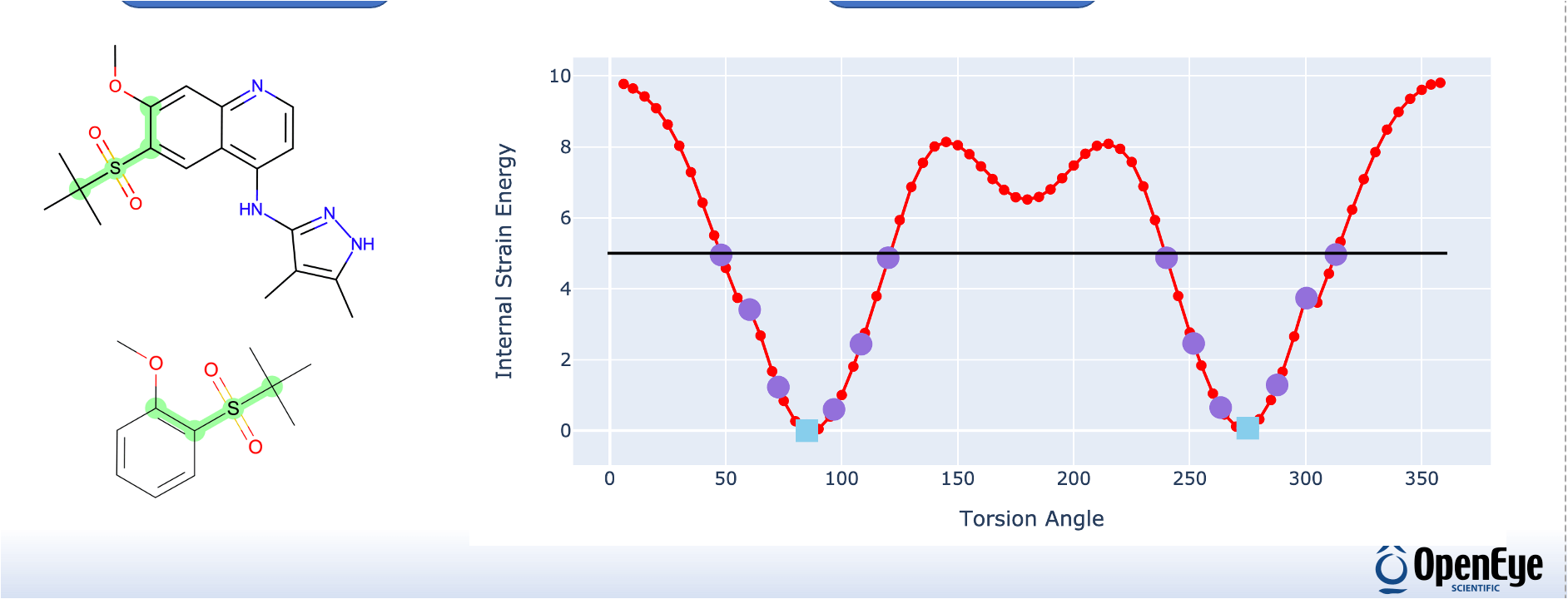
Quantum Mechanics
Gain valuable molecular insight at the electronic level of detail. OpenEye’s Gaussian and Psi4 workflows enable you to easily use quantum methods with various molecular modeling and simulations techniques in solving your most challenging scientific problems.
Machine Learning and Molecule Explainer
Build machine learning models, Assess generated models, Visualize and understand the results – all in one environment. Different from other machine learning methods, OpenEye lets you build models, assess quality, and understand your results in a chemically intuitive way.
-
 Learn more
Learn moreLarge Scale Virtual Screening
Search billions of molecules across different chemical libraries. OpenEye delivers 2D and 3D virtual screening solutions at the scale you need, the speed you want, and the computing cost you desire.
-
 Learn more
Learn moreStructural Biology
Get more accurate results with careful preparation of experimental structures for downstream modeling applications. OpenEye provides automated tools that can generate high quality models, understand protein motions, uncover cryptic pockets, and streamline your protein preparation process.
-
 Learn more
Learn moreLead Discovery
Rapidly discover new, biologically active chemical matter as potential leads in the drug discovery programs. Scientists trust OpenEye solutions for lead identification that are proven, fast and reliable.
-
 Learn more
Learn moreLead Optimization
Fine-tune profiles of your preclinical lead candidates for higher likelihood of a successful progression into clinical development. OpenEye provides a wide range of cutting-edge science to optimize both the biological activity and the properties of your lead series.
-
 Learn more
Learn moreAntibody Discovery
Rapidly analyze your antibody-based sequencing dataset with cloud computing. OpenEye provides you the ability to process tens of millions of sequences from the convenience of your web browser. Increase the number of your clonotype leads 5-10x compared to random colony screening.
-
 Learn more
Learn moreDrug Formulation
Leverage computational advances to cost-effectively predict downstream development profile of your drug leads. From pharmaceutical crystals predictions to intrinsic properties calculations, OpenEye provides you with the necessary tools to get ahead of your competitors.
-
 Learn more
Learn moreQuantum Mechanics
Gain valuable molecular insight at the electronic level of detail. OpenEye’s Gaussian and Psi4 workflows enable you to easily use quantum methods with various molecular modeling and simulations techniques in solving your most challenging scientific problems.
-
 Learn more
Learn moreMachine Learning and Molecule Explainer
Build machine learning models, Assess generated models, Visualize and understand the results – all in one environment. Different from other machine learning methods, OpenEye lets you build models, assess quality, and understand your results in a chemically intuitive way.
Trusted Science. Delivered the Way You Need.


