OMEGA
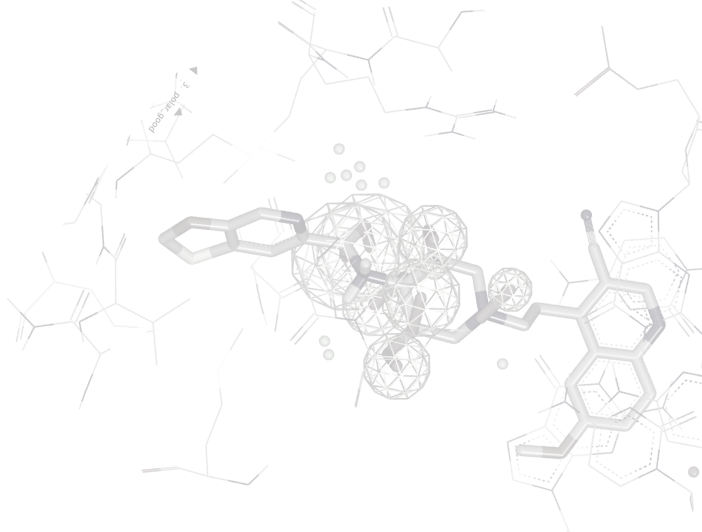
OMEGA is the most widely used, most highly cited conformation generation tool available. It robustly samples conformational space at high speed, using two different algorithms, one for broadly drug-like molecules and one for macrocyclic or highly flexible linear molecules.
OMEGA is very effective at reproducing bioactive conformations [1,2], and provides an optimal balance between speed and performance when used on large compound databases [3].
OMEGA conformational databases can be used as input to a variety of applications including docking engines (FRED), shape comparison tools (ROCS®), ligand posing (POSIT), and pharmacophore perception algorithms.
When compared to other tools OMEGA demonstrates both higher accuracy [4] and higher speed [5,6]. For more detailed information on OMEGA, check out the link below:
Documentation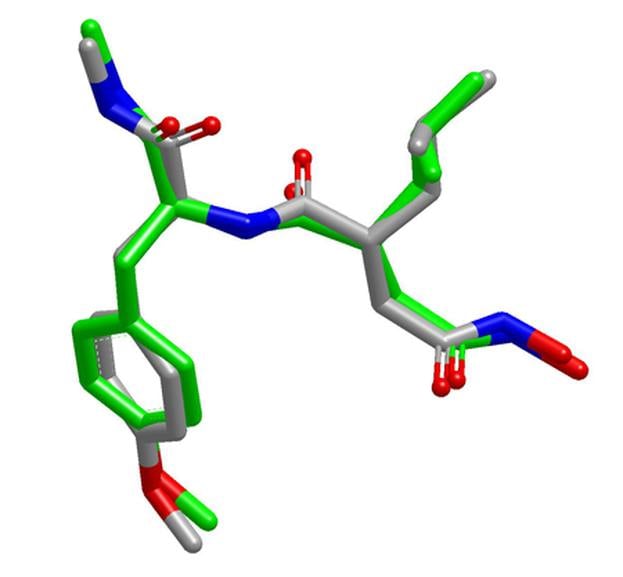
Features
- Flexible sampling methods. Torsion-driving with exhaustive and thompson sampling for drug-like molecules and distance geometry for macrocycles or very flexible linear molecules
- Very rapid (0.08 sec/molecule) rule-based sampling
- Diverse ensemble selection based on RMS deviation and strain energy
- User-configurable search resolution
- Automatic superposition of structural features
- Excellent reproduction of solid-state and solution conformations of drug-like molecules and macrocycles
- Distributed processing via MPI for all supported platforms
Learn more
OpenEye's OMEGA produces high-quality ensembles of bioactive conformations with the fastest speed when compared to other commercially available conformer generators for drug-like molecule.
For science details, WATCH OpenEye’s miniWebinar recording from August 2023 on OMEGA by Paul Hawkins, Ph.D., Product Evangelist, OpenEye, Cadence Molecular Sciences.
References
- Conformer Generation with OMEGA: Algorithm and Validation Using High Quality Structures from the Protein Databank and Cambridge Structural Database, Hawkins, P.C.D., Skillman, A.G., Warren, G.L. Ellingson, B.A. and Stahl, M.T., J. Chem. Inf. Model., 2010, 50, 572-584.
- Conformer generation with OMEGA: Learning from the dataset and analysis of failures, Hawkins, P.C.D., Nicholls, A.N., J. Chem. Inf. Model., 2012, 52, 2919-2936.
- Conformational Analysis of Drug-like Molecules Bound to Proteins: An Extensive Study of Ligand Reorganization upon Binding, Perola, E. and Charifson, P.S., J. Med. Chem. 2004,47, 2499-2510.
- High-Quality Dataset of Protein-Bound Ligand Conformations and its Application to Benchmarking Conformer Ensemble Generators, Friedrich, N.-O., Meyder, A., de Bruyn Kops, C., Sommer, K., Fachsenberg, F., Rarey, M., Kirchmair, J. J. Chem. Inf. Model. 2017, 57, 529-539.
- Benchmarking Conformer Ensemble Generators, Friedrich, N.-O. de Bruyn Kops, C. Fachsenberg, F. Sommer, K., Rarey, M. Kirchmair, J. J. Chem. Inf. Model. 2017, 57, 2719-2728.
- Decisions with Confidence: Application to the Conformation Sampling of Molecules in the Solid State, Hawkins, P.C.D., Wlodek, S., J. Chem. Inf. Model., 2020, 60, 3518-3533. (doi: 10.1021/acs.jcim.0c00358)
Webinar: Own Your Own Target with Target X
Webinar: Modular Molecular Modeling
Webinar: ML-Enabled integration of affinity prediction and lead discovery: 3D-QSAR
Webinar: Novel Hits from Beyond the Known: ROCS X
Webinar: Exploring the Uncharted: Discovery at Trillion-Scale with ROCS X
Resources
View Our Recent Webinars
Upcoming Webinar
Webinar: Target X: An Unobstructed View of Pockets
On Demand Webinars
Webinar: Own Your Own Target with Target X


