pKa Database
The protonation state of ionizable groups can play a critical role in understanding many of the biological processes involved in medicinal chemistry, such as membrane permeability, metabolism, and structure-activity relationships. While many programs attempt to predict pKas through parameterized models, large-scale access to primary measurements has previously been a difficult and tedious endeavor. pKa Prospector addresses this problem by providing rapid access to a comprehensive and relevant database of well-curated high-quality experimental pKa measurements. This database can be searched to identify the best data on the likely protonation state of novel molecules in biological systems.
Features
- A well-curated database of high-quality experimental pKa measurements
- Complete experimental data and reference for each measurement
- Search using rooted maximum common substructure (MCS)
- Sketch input structures
- Advanced filters and searching by
fingerprint similarity and by substructure - Auto-identification of ionizable groups
- Automated interface for finding relevant model compounds
- Highlighting simplifies identification of model compounds with multiple ionizable groups
- Retains history of previous searches for easy access
- Output results in spreadsheet form
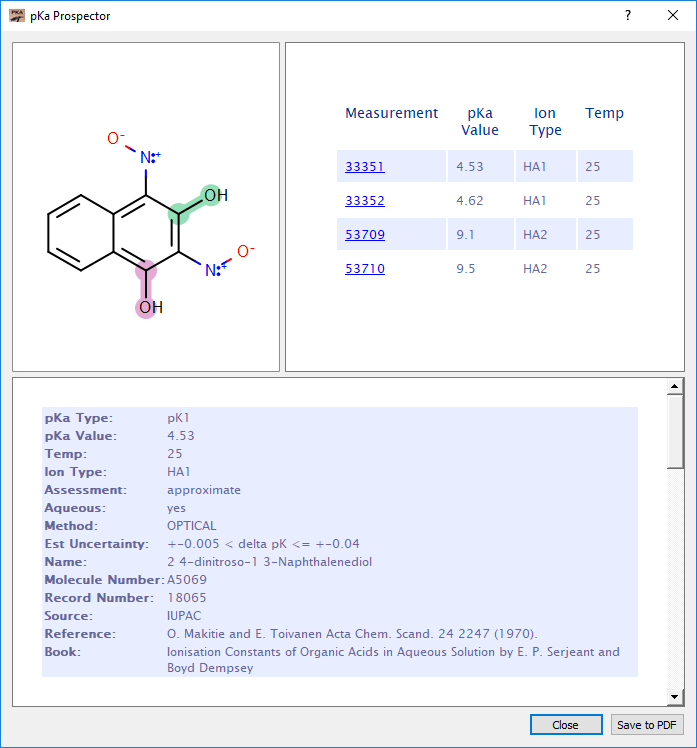
The built-in experimental pKa database was compiled by Tony Slater of pKaData Limited from a collection of IUPAC sources. Each measurement has been individually verified, curated, and assigned a metric of quality. There are more than 30,000 experiments across 12,000 molecules represented. The database is particularly relevant for medicinal chemistry due to the strong preponderance of room temperature aqueous measurements, the many molecules with multiple experimental records, and the presence of over three hundred different heterocycles.
In addition to the provided database, pKa Prospector provides the necessary tools to process and incorporate additional pKa measurements into the application. These measurements, whether from an internal database or external publication, are completely integrated into the search process and results viewer.
pKa Prospector uses a sophisticated, electronically-aware search method that can rapidly identify the most appropriate model compounds from the pool of experimental measurements. These high-quality search results, as opposed to a single context-free prediction from a trained model, provides the user with model compounds upon which to make pKa estimates based on the abundance or paucity of relevant primary data.

Experimental pKa Data
These pKa databases represent the extremely careful conversion of IUPAC's extensive compilations of experimental pKa values of organic acids and bases (in aqueous solution) from book form into fully curated computer-readable data, searchable by substructure.
Data Sources
- Base 1 (3775 molecules, 8766 pKas) Dissociation Constants of Organic Bases in Aqueous Solution, by D.D. Perin
- Acid 1 (1063 molecules, 2893 pKas) Dissociation Constants of Organic Acids in Aqueous Solution, by G. Kortum, W. Vogel and K. Andrussow
- Base 2 (4275 molecules, 7844 pKas) Dissociation Constants of Organic Bases in Aqueous Solution, Supplement 1972, by D.D. Perin
- Acid 2 (4584 molecules, 10912 pKas) Ionisation Constants of Organic Acids in Aqueous Solution, by E.P. Serjeant and Boyd Dempsey
The actual conversion process was done entirely by hand by Tony Slater of pKaData Limited to ensure accuracy and consistency. Tautomer enumeration was done using QUACPAC.
"The critical assessment of data quality is one of the major features of this seminal group of pKa compilations for weak organic acids and bases sponsored by the International Union of Pure and Applied Chemistry (IUPAC)"
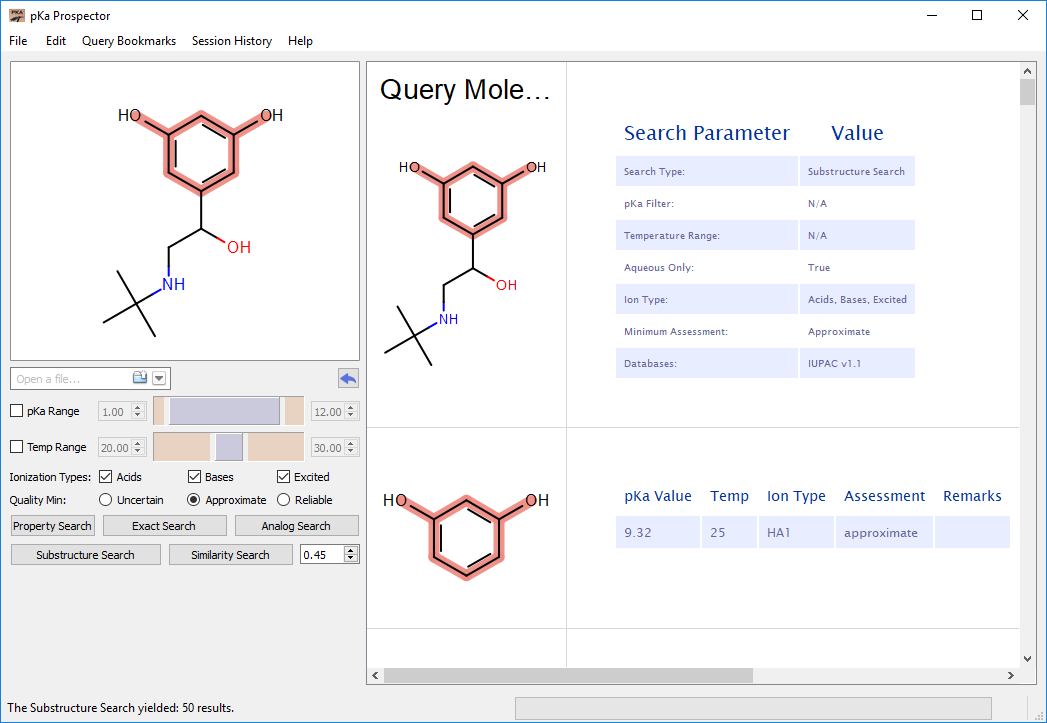
Features
The databases are delivered in both Microsoft Excel and CSV formats and can easily be merged with existing in-house databases with the IUPAC sourced data clearly identified.
- Molecule names and structures converted to SMILES
- IUPAC critical data quality assessment
- Data assigned to separate fields, e.g. ionic strength, concentration, and temperature
- Associated alphabetic data placed in separate field to numerical data (e.g. <5.3 assigned to two data fields) for enhanced search capabilities
- Full reference and method description for each record
- Ionization assignment for logD calculations
- Output results in spreadsheet form
Webinar: Own Your Own Target with Target X
Webinar: Modular Molecular Modeling
Webinar: ML-Enabled integration of affinity prediction and lead discovery: 3D-QSAR
Webinar: Novel Hits from Beyond the Known: ROCS X
Webinar: Exploring the Uncharted: Discovery at Trillion-Scale with ROCS X
Resources
View Our Recent Webinars
Upcoming Webinar
Webinar: Target X: An Unobstructed View of Pockets
Blog Post
Webinar: Own Your Own Target with Target X


