FRAGMENT REPLACEMENT & MOLECULAR DESIGN
BROOD
BROOD is designed to help explore chemical and property space around a hit or lead molecule. BROOD generates analogs of the lead by replacing selected fragments in the molecule with fragments that have similar shape and electrostatics, yet with selectively modified molecular properties.
BROOD fragment searching has multiple applications, including lead-hopping, side-chain enumeration, patent breaking, fragment merging, property manipulation, and patent protection by SAR expansion.
Using OpenEye’s BROOD software for scaffold hopping, we successfully discovered a new chemical series from our lead compound, with equipotency and improved stability. – Laurent D., Lundbeck
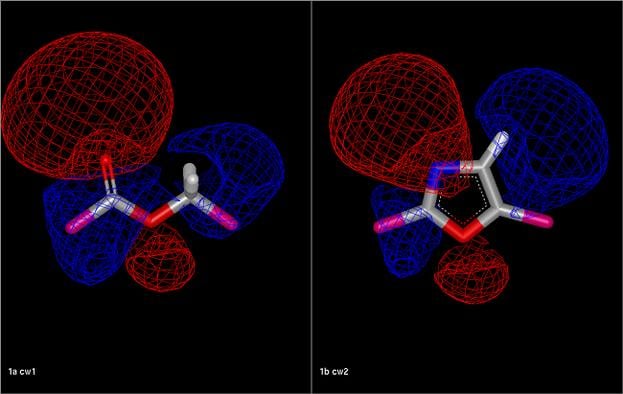
Comparison of an ester fragment and an oxazole fragment showing the electrostatic isopotential contour surfaces. The electrostatic Tanimoto between the two fragments is 0.54.
Features
- Graphical query editing including the ability to easily modify the similarity force-field as well as add and remove chemical constraints such as those known to be critical to the SAR under development
- Graphical interface to facilitate physical property analysis and real-time property filtering of millions of potential molecules
- Integrated estimation of synthetic accessibility
- Construction and assessment of new molecule series in a protein active-site
- Hierarchical organization of the hitlist of analog molecules along with a new graphical interface designed specifically to explore and edit the hitlists
- Database of more than 4 million medicinally relevant fragments provided and utilities to aid users in augmenting this database with novel fragments from their corporate collections
- Tools for modelers and chemists to communicate their ideas and preferences including favorite molecules list management, molecular annotation, view bookmarking and hitlist subsetting
RESOURCES
