OpenEye Scientific is pleased to announce the release of OpenEye Applications and Toolkits 2020.2, which contains several key upgrades and improved performance.
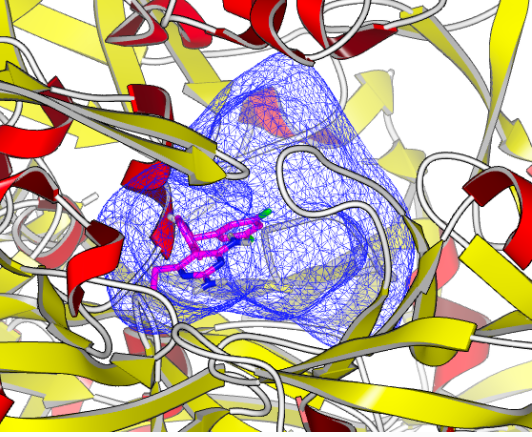
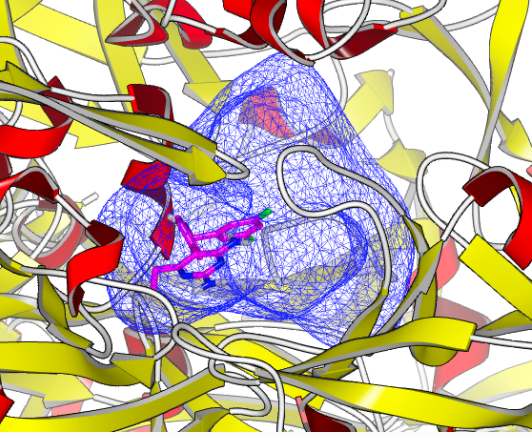
OEDocking receptor, along with the target structure and the bound ligand, for the 2IKO Human Renin Complexed with Inhibitor.
HIGHLIGHTS
- New Fragment Library created for OMEGA
Designed to reduce the run-time of large-scale conformer generation, the enhanced fragment library, consisting of more than 500,000 fragments, shows a dramatic increase in OMEGA run-time performance while generating the same high-quality conformers as the previous version.
Read more about this highlight here. - Improved Receptors in OEDOCKING
Receptors are now an integral part of the Design units, taking full advantage of the protein structures prepared using SPRUCE. This feature facilitates the easier use of docked and posed structures in downstream modeling applications.
Read more about this highlight here. - New Protein Force Field in SZYBKI
The AMBER FF14SB protein force field, the most widely used protein force field for molecular dynamics or any other force-field based calculations, now is available in SZYBKI for optimizing both protein and protein-ligand complexes.
Read more about this highlight here.
These application packages include the usual support for Linux, macOS, and Windows. In addition, the toolkit libraries include the usual support for C++, C#, Java, and Python.
For more details and additional key updates to the 2020.2 Applications and Toolkits release, please read the full Application and Toolkit release notes.
To learn more about the highlights and updates in the latest release, please join us for a webinar "Applications and Toolkits 2020.2 Update," with Shyamal Nath, PhD, Head of Molecular Modeling at OpenEye on Thursday 21 January 2021, at the following times:
- 4 - 5 p.m. | Central European Time (CET)
- 3 - 4 p.m. | Greenwich Mean Time (GMT)
- 10 - 11 a.m. | U.S. Eastern Standard Time (EST)
- 9 - 10 a.m. | U.S. Central Standard Time (CST)
- 8 - 9 a.m. | U.S. Mountain Standard Time (MST)
- 7 - 8 a.m. | U.S. Pacific Standard Time (PST)
Please register for the webinar, which will be hosted through GoToWebinar.
For Asia customers or for those who are unable to attend the “live” webinar, a recording of the webinar also will be available. To receive the link to the recorded webinar, please register for the webinar.
We hope you can attend.