GENERATING AND RENDERING STRUCTURES
OEDepict TK
OEDepict TK offers rapid 2D coordinate generation and artful rendering of chemical structures. 2D coordinates can be generated from connection tables (e.g. SMILES) or 3D structures. Multiple image file formats are supported including:
- PNG (Portable Network Graphics)
- PDF (Portable Document Format)
- SVG (Scalable Vector Graphics)
- Postscript
- Encapsulated PostScript
.jpg?width=623&height=415&name=oedepiction%20(1).jpg)
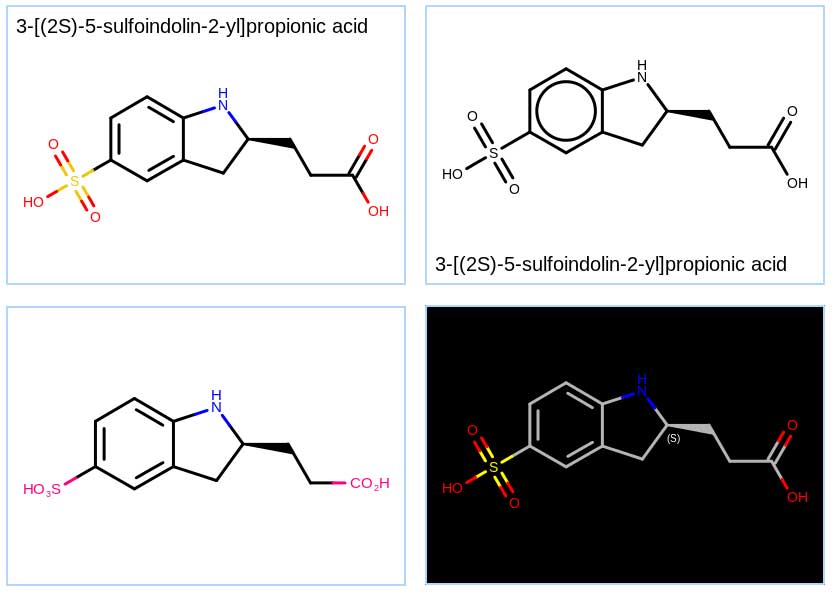
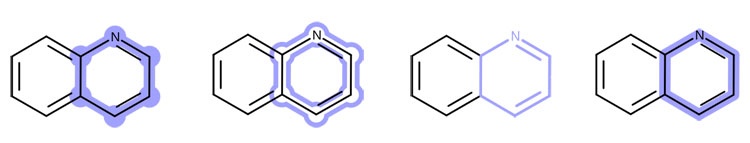
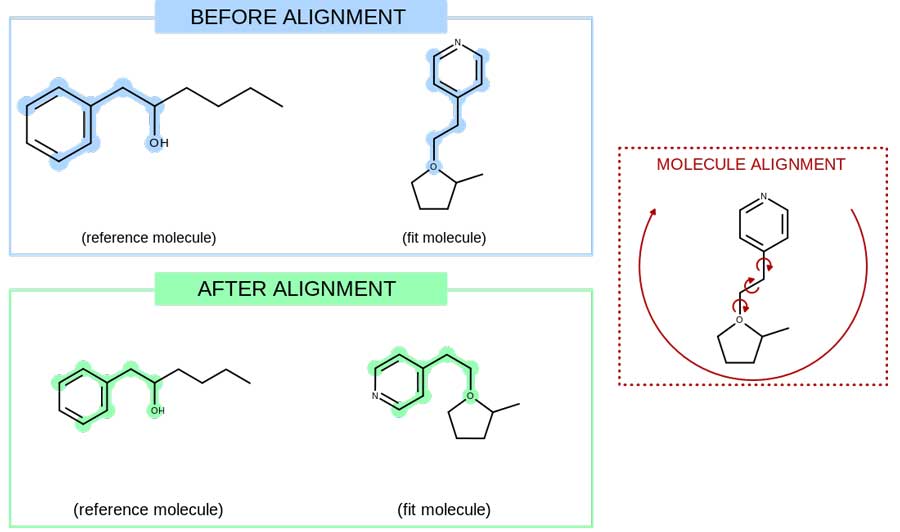
In addition to standard depiction routines, OEDepict TK provides a powerful graphics engine which supports:
- Basic shape drawing
- Highly customizable molecule depiction
- Variety of highlighting styles
- Customizable layout options (including grids and multi-page formatting)
- Depictions aligned by MCS or substructure
- MDL Query and MDL Reaction depictions
For more detailed information on OEDepict TK, check out the link below:
DocumentationModeling
The Modeling suite of toolkits provides the core functionality underlying OpenEye's defining principle that shape & electrostatics are the two fundamental descriptors determining intermolecular interactions. Many of the toolkits in the Modeling suite are directly associated with specific OpenEye applications and can, therefore, be used to create new or extend existing functionality associated with those applications.
- OEChem TK Core chemistry handling and representation as well as molecule file I/O
- FastROCS™ TK Real-time shape similarity for virtual screening, lead hopping & shape clustering
- OEDocking TK Molecular docking and scoring
- Omega TK Conformer generation
- Shape TK 3D shape description, optimization, and overlap
- SiteHopper TK Rapid comparison of protein binding sites
- Spicoli TK Surface generation, manipulation, and interrogation
- Spruce TK Protein preparation and modeling
- Szybki TK Force field based focused optimization and entropy estimations
- OEFF TK Force fields and general optimization tools
- Szmap TK Understanding water interactions in a binding site
- Zap TK Calculate Poisson-Boltzmann electrostatic potentials
- Bioisostere TK 3D fragment similarity, lead optimization, and analog generation
- Eon TK Molecular shape and electrostatic overlap
Cheminformatics
The Cheminformatics suite of toolkits provides the core foundation upon which all the OpenEye applications and remaining toolkits are built.
- OEChem TK Core chemistry handling and representation as well as molecule file I/O
- OEDepict TK 2D Molecule rendering and depiction
- Grapheme™ TK Advanced molecule rendering and report generation
- GraphSim TK 2D molecular similarity (e.g. fingerprints)
- Lexichem™ TK Name-to-structure, structure-to-name, foreign language translation
- MolProp TK Molecular property calculation and filtering
- Quacpac TK Tautomer enumeration and charge assignment
- OEMedChem TK Matched molecular pair analysis, fragmentation utilities, and molecular complexity metrics
- Saiph TK: Extracting, transforming, and loading (ETL) files and records
Webinar: OpenEye's Free energy prediction for drug discovery: Ideas at breakfast, discoveries by lunch
Webinar: Own Your Own Target with Target X
miniCUP Basel 2026
Cadence Tool Reveals Druggable Sites with Over 90% Accuracy
Resources
Glimpse the Future through News, Events, Webinars and more
News
ROCS X: AI-Enabled Molecular Search Unlocks Trillions
Webinar
Webinar: OpenEye's Free energy prediction for drug discovery: Ideas at breakfast, discoveries by lunch


