Shape TK
Shape TK is the basis for the ROCS® application for shape-based similarity searching. The toolkit allows for fine control over optimization methods, molecule treatment and the nature of the query (grids, generic shapes). Shape TK facilitates the calculation of molecular descriptors for shape (steric multipoles), volume overlap between molecules, and spatial similarity of chemical groups (color force field), as well as the optimization of the latter two quantities.
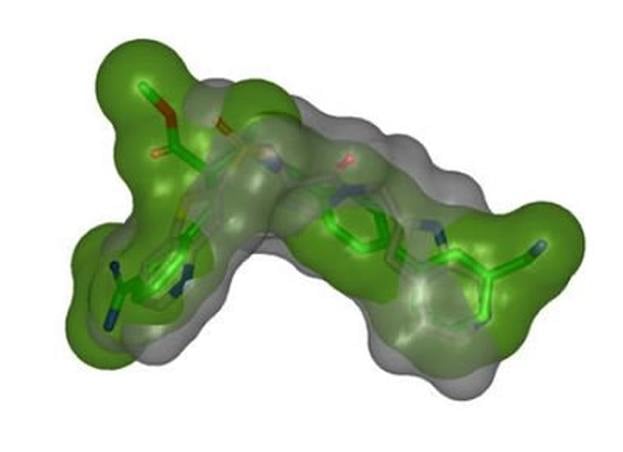

Shape TK - a flexible toolkit for shape and color overlay
Shape TK features include shape and color overlay, optimized overlays for queries composed solely of color atoms or Gaussians, and support for using grids as database entities for overlay.
Examples of use include real-space fitting to electron density, shape fingerprinting [1], discrimination between agonists and antagonists based on shared shapes [2], generation of composite queries from solvent mapping [3], and aligning and comparing protein active sites [4].
For more detailed information on Shape TK, check out the link below:
Modeling
The Modeling suite of toolkits provides the core functionality underlying OpenEye's defining principle that shape & electrostatics are the two fundamental descriptors determining intermolecular interactions. Many of the toolkits in the Modeling suite are directly associated with specific OpenEye applications and can, therefore, be used to create new or extend existing functionality associated with those applications.
- OEChem TK Core chemistry handling and representation as well as molecule file I/O
- FastROCS™ TK Real-time shape similarity for virtual screening, lead hopping & shape clustering
- OEDocking TK Molecular docking and scoring
- Omega TK Conformer generation
- Shape TK 3D shape description, optimization, and overlap
- SiteHopper TK Rapid comparison of protein binding sites
- Spicoli TK Surface generation, manipulation, and interrogation
- Spruce TK Protein preparation and modeling
- Szybki TK Force field based focused optimization and entropy estimations
- OEFF TK Force fields and general optimization tools
- Szmap TK Understanding water interactions in a binding site
- Zap TK Calculate Poisson-Boltzmann electrostatic potentials
- Bioisostere TK 3D fragment similarity, lead optimization, and analog generation
- Eon TK Molecular shape and electrostatic overlap
Cheminformatics
The Cheminformatics suite of toolkits provides the core foundation upon which all the OpenEye applications and remaining toolkits are built.
- OEChem TK Core chemistry handling and representation as well as molecule file I/O
- OEDepict TK 2D Molecule rendering and depiction
- Grapheme™ TK Advanced molecule rendering and report generation
- GraphSim TK 2D molecular similarity (e.g. fingerprints)
- Lexichem™ TK Name-to-structure, structure-to-name, foreign language translation
- MolProp TK Molecular property calculation and filtering
- Quacpac TK Tautomer enumeration and charge assignment
- OEMedChem TK Matched molecular pair analysis, fragmentation utilities, and molecular complexity metrics
- Saiph TK: Extracting, transforming, and loading (ETL) files and records
References
-
Small molecule shape-fingerprints Haigh, J. A., Pickup, B. T., Grant, J. A., Nicholls, A., J. Chem. Inf. Model., 2005, 45, 673.
-
Scaffold hopping, synthesis and structure-activity relationships of 5,6-diaryl-pyrazine-2-amide derivatives: a novel series of CB1 receptor antagonists Boström, J., Berggren, K., Elebring, T., Greasley, P.J., Wilstermann, M., Bioorg Med Chem., 2007, 15, 4077.
-
The use of fake ligands from computational solvent mapping in ligand and structure-based virtual screening Hall, D. R., Enyedy, I., Future Med Chem. 2016 Oct;8(15):1815-1823. doi: 10.4155/fmc-2016-0115. Epub 2016 Sep 15. PMID: 27630057.
-
SiteHopper - a unique tool for binding site comparison
Webinar: OpenEye's Free energy prediction for drug discovery: Ideas at breakfast, discoveries by lunch
Webinar: Own Your Own Target with Target X
miniCUP Basel 2026
Cadence Tool Reveals Druggable Sites with Over 90% Accuracy
Resources
Glimpse the Future through News, Events, Webinars and more
News
ROCS X: AI-Enabled Molecular Search Unlocks Trillions
Webinar
Webinar: OpenEye's Free energy prediction for drug discovery: Ideas at breakfast, discoveries by lunch


