OpenEye is pleased to announce the release of OpenEye Toolkits v2019.Oct. These libraries include the usual support for C++, C#, Java, and Python.
HIGHLIGHTS
- Spruce TK, a new toolkit for preparing biomolecular structures for modeling applications, is now available in both C++ and Python APIs.
- OEChem TK now provides improved substructure search capability.
- SMIRNOFF, a small molecule force field from the Open Force Field Initiative, is now integrated into OEFF TK.
Spruce TK: A new toolkit for biomolecular structure preparation
Proteins prepared in a particular state are essential as input to many modeling applications, such as docking with FRED or HYBRID, pose prediction with POSIT, identifying favorable and unfavorable waters with SZMAP or free energy calculations using molecular dynamics simulations. The 2019.Oct release introduces Spruce TK, which generates such protein structures from experimental biomolecular structure data.
Users can easily prepare proteins using either C++ or Python APIs (see OEMakeDesignUnits), where the output of the file is the OEDesignUnit class and is serialized to a new OEDU file format. The prepared system is componentized into molecular categories including protein, ligand, nucleic, solvent, and many other molecular types common to biomolecular experiments. By default, hydrogens are added to systems using a tautomer search for any ligand, cofactor, or other non-standard residue molecules. In addition, the system is split into distinct biological units, and all final design unit structures are superposed into the common frame of reference of a parent structure. For more details, see Spruce theory.
Spruce TK has been tested across a broad range of industry-relevant targets. It has been used in virtual screening and molecular dynamics simulations in Orion, as well as to prepare structures for loading into the MacroMolecular Data Service (MMDS).
OEChem TK: Fast Substructure Search
Substructure search capability in OEChem TK has been improved, allowing users to search tens of millions of molecules in seconds. OEChem TK uses a screen-based approach to accelerate the search process. Screens – bit vectors encoding global and local features of molecules – are prebuilt and can be used to rapidly eliminate structures that could not be matched to a query molecule. Only molecules that pass the screen are subjected to the computationally more expensive atom-by-atom matching for validation. OEChem TK provides three types of built-in screens based on the source of the query molecule: MDL, SMARTS, and Molecule. All screens are rigorously tested using diverse sets of queries and multiple molecular databases to ensure that they do not eliminate true positive matches.
The new multi-threaded substructure search can be performed in two modes: in-memory and molecule-database. The in-memory mode provides the fastest way to search a dataset, but it is memory-intensive as it holds both screens and molecules in memory. The molecule-database mode keeps only the screens in memory; only molecules with unmatching screens have to be loaded into memory. This method can be slower but uses significantly less memory, allowing users to search larger datasets.
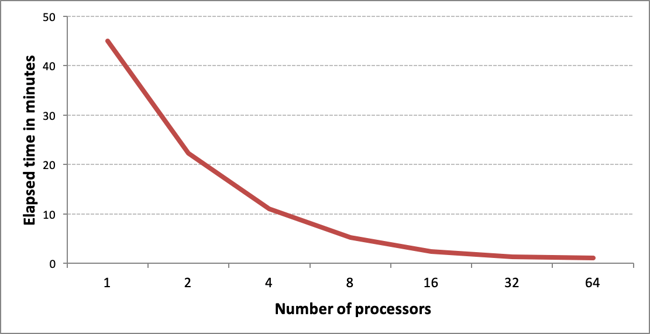
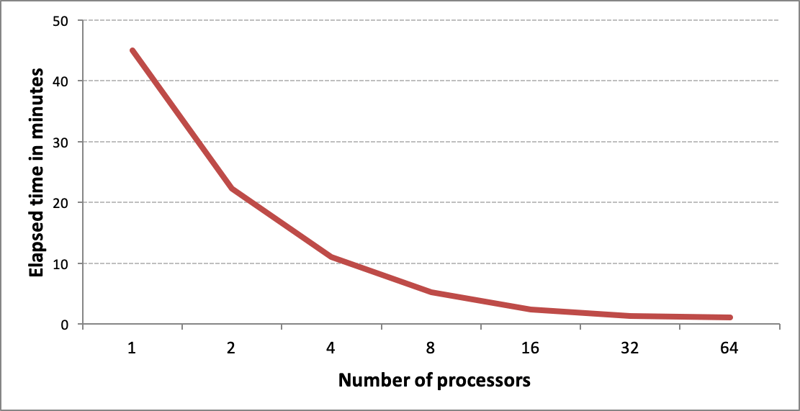
Multi-threaded screen generation performance for 1M MDL 896-bit long screens benchmarked on an r5.16xlarge AWS instance
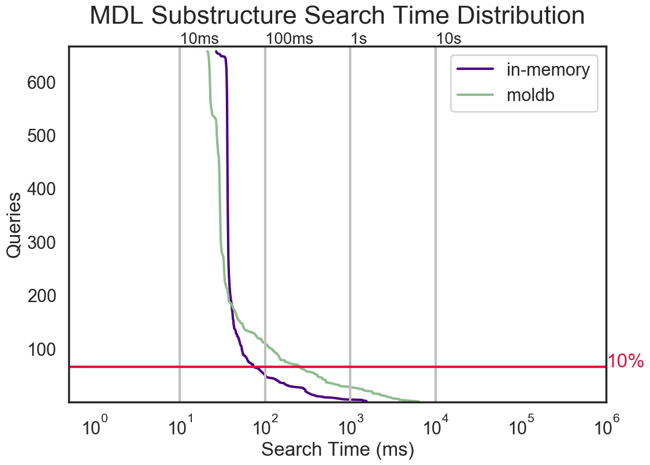
Multi-threaded substructure search performance for an in-house diverse set of 650 MDL queries against an Enamine REAL Diverse 24.1M database benchmarked on an r5.16xlarge AWS instance
See the Molecule searching chapter of OEChem TK for examples using the new OESubSearchDatabase class. For the full list of new preliminary APIs, see OEChem TK 2.3.0.
OEFF TK: SMIRNOFF
The 2019.Oct release incorporates the SMIRks Native Open Force Field (SMIRNOFF), a small molecule force field from the Open Force Field Initiative, into OEFF TK. The force field can handle almost all pharmaceutically relevant chemical space. SMIRNOFF differs from traditional force fields mainly in the data representation of the chemistry. It uses extended SMARTS strings for parameters instead of atom types, greatly reducing the number of interdependent parameters. It currently implements the functional form of the AMBER force field, but the Open Force Field Initiative is actively examining improvements to this. The most up-to-date versions of SMIRNOFF will be included in OEFF TK whenever possible.
GENERAL NOTICES
- A new search engine that provides better hits and shows results in a cleaner format has been added to the documents.
- The 2019.Oct release now uses SWIG 3.0.12 for generating C#, Java, and Python wrappers. C# and Python use SWIG’s built-in support for wrapping C++ std::vector and string types.
PLATFORM SUPPORT
- This is the last release to support macOS 10.12. Full support for macOS 10.15 will be added in the next release.
- This is the last release to support RHEL6. Full support for RHEL8 will be added in the next release.
- The 2019.Oct release skips Java support for macOS. For Java support on RHEL6, please contact support@eyesopen.com.
AVAILABILITY
OpenEye Toolkits v2019.Oct are now available for download. Existing licenses will continue to work, but if a new license is required, please contact your account manager or email sales@eyesopen.com.
See the Release Notes for full and specific details on improvements and fixes.
RESOURCES
|
• Download |
• OEChem TK |
About OpenEye Scientific
OpenEye Scientific is a privately held company headquartered in Santa Fe, New Mexico, with offices in Boston, Cologne, and Tokyo. It was founded in 1997 to develop large-scale molecular modeling applications and toolkits. Primarily aimed towards drug discovery and design, areas of application include:
- Cheminformatics
- Structure Generation
- Shape Comparison
- Docking
- Fragment Replacement
- Electrostatics
- Crystallography
- Visualization
The software is designed for scientific rigor as well as speed, scalability, and platform independence. OpenEye makes most of its technology available as toolkits—programming libraries suitable for custom development. OpenEye software typically is distributable across multiple processors and runs on Linux, Windows, and macOS.
For additional information
Jeffrey Grandy
VP Sales
+1-415-863-3032
Email: sales@eyesopen.com
For additional information
Jeffrey Grandy
VP Sales
+1-415-863-3032
Email: sales@eyesopen.com