OpenEye is pleased to announce the release of OpenEye Toolkits v2016.Oct. These libraries include the usual support for C++, Python, C#, and Java.
NEW FEATURES
- Protein Preparation: New hydrogen placement functionality has been added to the protein preparation workflow.
- Protein-Ligand Interaction: Perception of potential protein-ligand interactions, intramolecular interactions, and missing potential interactions has been enhanced.
- CUDA Implementation in FastROCS TK: The internal GPU implementation of FastROCS now uses CUDA.
Protein Preparation
OEBio TK now provides a new protein preparation function, OEPlaceHydrogens. This function adds and optimizes the positions of hydrogens in a protein to generate optimum hydrogen bonding networks while avoiding atom clashes. In addition, it can flip functional groups that are sometimes modeled incorrectly, such as histidine, asparagine, and glutamine sidechains.
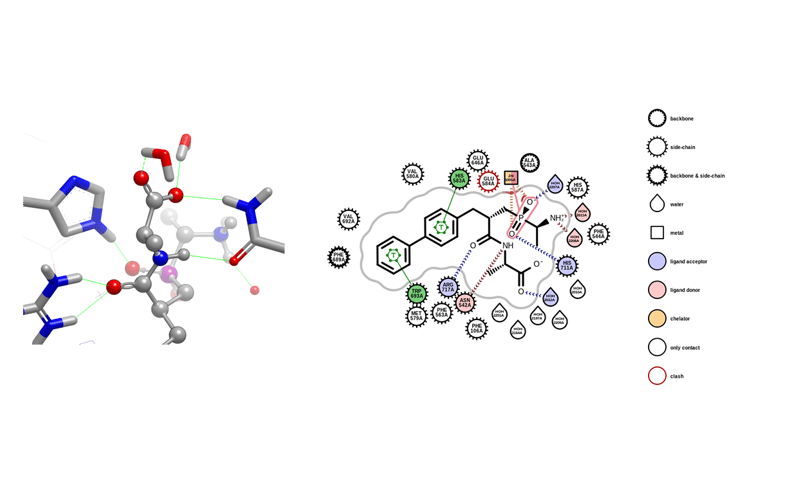
OEPlaceHydrogens can be applied broadly to many protein-ligand structures. It uses SMARTS patterns rather than relying on naming conventions. A new example program, proteinprep, demonstrates this functionality. This release debuts this functionality; it is still under active development.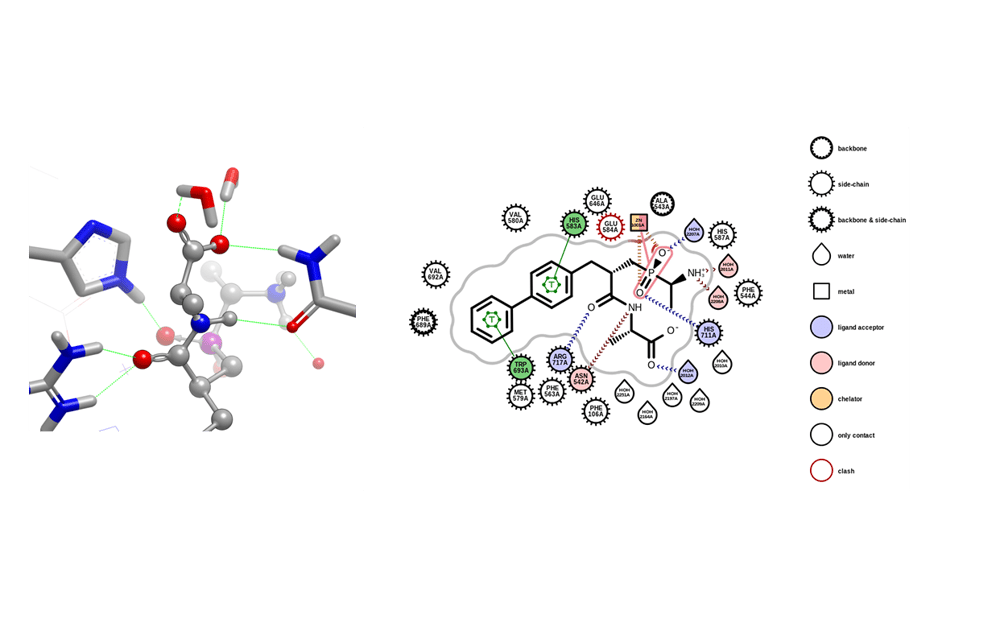
An example of a hydrogen bonding network in 1R1H along with the corresponding protein-ligand map
Protein-Ligand Interaction
Significant improvements to the perception of potential protein-ligand interactions, intramolecular interactions, and missing potential interactions in OEBio TK have been made. A new type of interaction, halogen bond, joins hydrogen bond, salt bridge, chelation, and pi- and T-aromatic stacking in the set of perceived interaction types. These interactions provide qualitative hints about biological interactions that may be important for binding, and can be helpful for visualizing interactions and clashes.
An interface for modifying the geometric definitions for each interaction has also been introduced. While the default bond geometries have been carefully constructed using literature data, users can also customize the interaction parameters.
CUDA Implementation in FastROCS TK
The internal GPU implementation of FastROCS has been ported from OpenCL to CUDA. GPU computing with CUDA allows OpenEye to better support NVIDIA hardware and facilitates our ability to improve performance. The switch from OpenCL to CUDA does not, however, change the results returned by FastROCS and should not impact the deployment of FastROCS TK on customer machines.
PLATFORM SUPPORT
- The 2016.Oct release adds support for Ubuntu16 for C++, Java, Python 2.7, and Python 3.x.
- The 2016.Oct release is the last release to support OSX 10.9. The 2017.Feb release will add full support for MacOS Sierra 10.12.
Note: OpenEye is planning to phase out Python 2 support by the 2017.Oct release. As this is a substantial change for us and our customers, we are willing to help with code migration. Please contact support@eyesopen.com for more details.
AVAILABILITYOpenEye Toolkits v2016.Oct are now available for download. Existing licenses will continue to work, but if a new license is required, please contact your account manager or email sales@eyesopen.com.
See the release notes for full and specific details on improvements and fixes.
RESOURCES
|
About OpenEye Scientific Software
OpenEye Scientific Software Inc. is a privately held company headquartered in Santa Fe, NM, with offices in Boston, Cologne, and Tokyo. It was founded in 1997 to develop large-scale molecular modeling applications and toolkits. Primarily aimed towards drug discovery and design, areas of application include:
- Cheminformatics
- Structure Generation
- Shape Comparison
- Docking
- Fragment Replacement
- Electrostatics
- Crystallography
- Visualization
The software is designed for scientific rigor, as well as speed, scalability and platform independence. OpenEye makes most of its technology available as toolkits - programming libraries suitable for custom development. OpenEye software typically is distributable across multiple processors and runs on Linux, Windows and Mac OS X.
For additional information
Jeffrey Grandy
VP Sales
+1-415-863-3032
Email: sales@eyesopen.com