OpenEye is pleased to announce the release of OpenEye Toolkits v2018.Oct. These libraries include the usual support for C++, Python, C#, and Java.
HIGHLIGHTS
- Omega TK now includes a method specifically tuned to sample macrocyclic conformational space.
- FastROCS TK is now available in C++ and Java.
- Quacpac TK includes improvements to the tautomer functionality.
Omega TK: Macrocycle
A new distance geometry-based, conformational space-sampling method has been added to Omega TK as a preliminary API. The approach is based on a rewritten version of the original OMEGA distance geometry algorithm and was first introduced in the OMEGA v3.0.0 app. The distance geometry-based approach opens up a new avenue for exploring conformer ensembles in Omega TK.
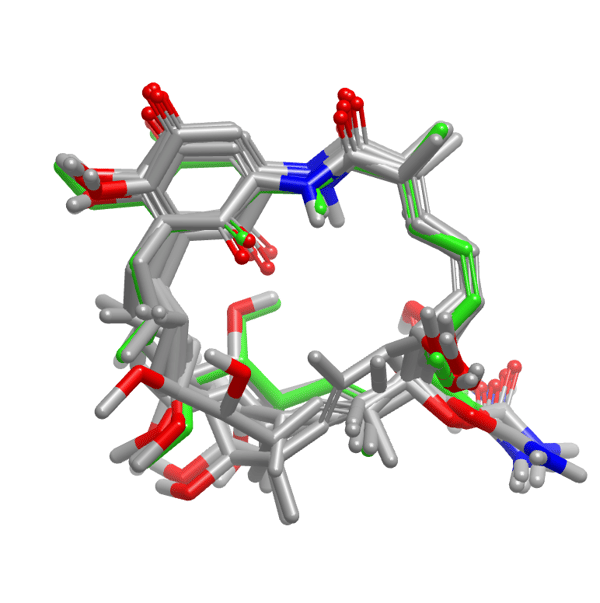
Conformer ensemble subset for geldanamycin, an inhibitor of heat shock protein 90, Hsp90 (PDB: 1YET), generated using the new macrocycle method.
FastROCS C++ and Java Support added
Previously, FastROCS TK was only available in Python. This release extends support to C++ and Java. All classes, functions, and constants are now available in C++, Java, and Python. Additionally, users running FastROCS TK on Tesla GPUs will see a 10 - 30% speed-up over the previous version, depending on the system.
Quacpac TK: Tautomers
This release adds several improvements to the tautomer functionality in Quacpac TK. Tautomer handling in molecular modeling and cheminformatics usually includes three important functions:
- Generating a canonical protomer representation for duplicate-free storage
- Generating a single tautomer suitable for visualization by chemists and modelers
- Generating an ensemble of aqueous-phase tautomers suitable for molecular modeling
Example of generating an ensemble of biologically relevant tautomers
This release provides a new API to cover each of these use-cases. The algorithm has also been improved to handle memory usage in cases where there are a large number of possible tautomeric forms.
GENERAL NOTICES
- The release cycle of OpenEye Toolkits has been changed. See the Release Cycle section for more information.
- FastROCS TK no longer supports Nvidia Tesla, GeForce, and Quadro cards with a Compute Capability older than 3.0. This includes all GPUs with Fermi architecture (e.g., Tesla C2050 and GeForce GTX 430). Users running one of the older GPUs who cannot upgrade their hardware should not update to this version.
- Spruce TK now requires a specific license. For information on Spruce TK licensing, please contact sales@eyesopen.com.
- Improvements have been made in the Python 3 toolkits with regard to wrapping optimization. This results in Python 3 performance being comparable to Python 2.
PLATFORM SUPPORT
- This is the last release to support GCC 4.4 on RHEL6. GCC 4.8 will be the oldest targeted compiler in future toolkit releases.
- This is the last release to support macOS 10.11.
- This release adds support for Ubuntu 18.
- This release adds support for Python 3.7 on macOS and Linux. On Windows, Python 3.7 will be supported in the next release.
AVAILABILITY
OpenEye Toolkits v2018.Oct are now available for download. Existing licenses will continue to work, but if a new license is required, please contact your account manager or email sales@eyesopen.com.
See the Release Notes for full and specific details on improvements and fixes.
RESOURCES
|
About OpenEye Scientific
OpenEye Scientific is a privately held company headquartered in Santa Fe, New Mexico, with offices in Boston, Cologne, and Tokyo. It was founded in 1997 to develop large-scale molecular modeling applications and toolkits. Primarily aimed towards drug discovery and design, areas of application include:
- Cheminformatics
- Structure Generation
- Shape Comparison
- Docking
- Fragment Replacement
- Electrostatics
- Crystallography
- Visualization
The software is designed for scientific rigor as well as speed, scalability, and platform independence. OpenEye makes most of its technology available as toolkits—programming libraries suitable for custom development. OpenEye software typically is distributable across multiple processors and runs on Linux, Windows, and macOS.
For additional information
Jeffrey Grandy
VP Sales
+1-415-863-3032
Email: sales@eyesopen.com

